#include <Marginals.h>
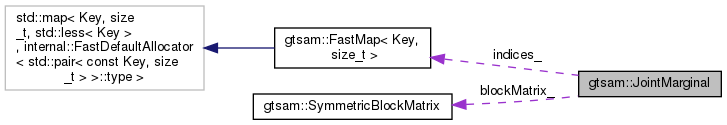
Collaboration diagram for gtsam::JointMarginal:
Public Member Functions | |
| JointMarginal () | |
| Default constructor only for wrappers. | |
| Matrix | operator() (Key iVariable, Key jVariable) const |
| Matrix | at (Key iVariable, Key jVariable) const |
| Matrix | fullMatrix () const |
| void | print (const std::string &s="", const KeyFormatter &formatter=DefaultKeyFormatter) const |
Protected Member Functions | |
| JointMarginal (const Matrix &fullMatrix, const std::vector< size_t > &dims, const KeyVector &keys) | |
Protected Attributes | |
| SymmetricBlockMatrix | blockMatrix_ |
| KeyVector | keys_ |
| FastMap< Key, size_t > | indices_ |
Friends | |
| class | Marginals |
Detailed Description
A class to store and access a joint marginal, returned from Marginals::jointMarginalCovariance and Marginals::jointMarginalInformation
Member Function Documentation
◆ at()
Synonym for operator()
◆ fullMatrix()
|
inline |
The full, dense covariance/information matrix of the joint marginal.
◆ operator()()
Access a block, corresponding to a pair of variables, of the joint marginal. Each block is accessed by its "vertical position", corresponding to the variable with nonlinear Key iVariable and "horizontal position", corresponding to the variable with nonlinear Key jVariable.
For example, if we have the joint marginal on a 2D pose "x3" and a 2D landmark "l2", then jointMarginal(Symbol('x',3), Symbol('l',2)) will return the 3x2 block of the joint covariance matrix corresponding to x3 and l2.
- Parameters
-
iVariable The nonlinear Key specifying the "vertical position" of the requested block jVariable The nonlinear Key specifying the "horizontal position" of the requested block
◆ print()
| void gtsam::JointMarginal::print | ( | const std::string & | s = "", |
| const KeyFormatter & | formatter = DefaultKeyFormatter |
||
| ) | const |
The documentation for this class was generated from the following file:
- /home/docs/checkouts/readthedocs.org/user_builds/gtsam-jlblanco-docs/checkouts/latest/gtsam/nonlinear/Marginals.h
 1.8.13
1.8.13