#include <GaussianFactorGraph.h>
Public Types | |
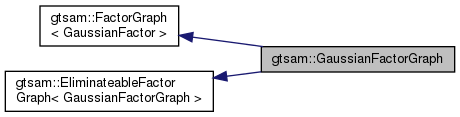
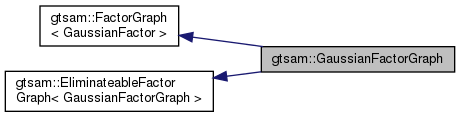
| typedef GaussianFactorGraph | This |
| Typedef to this class. | |
| typedef FactorGraph< GaussianFactor > | Base |
| Typedef to base factor graph type. | |
| typedef EliminateableFactorGraph< This > | BaseEliminateable |
| Typedef to base elimination class. | |
| typedef std::shared_ptr< This > | shared_ptr |
| shared_ptr to this class | |
| typedef KeySet | Keys |
| typedef GaussianFactor | FactorType |
| factor type | |
| typedef std::shared_ptr< GaussianFactor > | sharedFactor |
| Shared pointer to a factor. | |
| typedef sharedFactor | value_type |
| typedef FastVector< sharedFactor >::iterator | iterator |
| typedef FastVector< sharedFactor >::const_iterator | const_iterator |
| typedef EliminationTraits< FactorGraphType > | EliminationTraitsType |
| Typedef to the specific EliminationTraits for this graph. | |
| typedef EliminationTraitsType::ConditionalType | ConditionalType |
| Conditional type stored in the Bayes net produced by elimination. | |
| typedef EliminationTraitsType::BayesNetType | BayesNetType |
| Bayes net type produced by sequential elimination. | |
| typedef EliminationTraitsType::EliminationTreeType | EliminationTreeType |
| Elimination tree type that can do sequential elimination of this graph. | |
| typedef EliminationTraitsType::BayesTreeType | BayesTreeType |
| Bayes tree type produced by multifrontal elimination. | |
| typedef EliminationTraitsType::JunctionTreeType | JunctionTreeType |
| Junction tree type that can do multifrontal elimination of this graph. | |
| typedef std::pair< std::shared_ptr< ConditionalType >, std::shared_ptr< _FactorType > > | EliminationResult |
| typedef std::function< EliminationResult(const FactorGraphType &, const Ordering &)> | Eliminate |
| The function type that does a single dense elimination step on a subgraph. | |
| typedef std::optional< std::reference_wrapper< const VariableIndex > > | OptionalVariableIndex |
| typedef std::optional< Ordering::OrderingType > | OptionalOrderingType |
| Typedef for an optional ordering type. | |
Public Member Functions | |
| void | add (const GaussianFactor &factor) |
| void | add (const sharedFactor &factor) |
| void | add (const Vector &b) |
| void | add (Key key1, const Matrix &A1, const Vector &b, const SharedDiagonal &model=SharedDiagonal()) |
| void | add (Key key1, const Matrix &A1, Key key2, const Matrix &A2, const Vector &b, const SharedDiagonal &model=SharedDiagonal()) |
| void | add (Key key1, const Matrix &A1, Key key2, const Matrix &A2, Key key3, const Matrix &A3, const Vector &b, const SharedDiagonal &model=SharedDiagonal()) |
| template<class TERMS > | |
| void | add (const TERMS &terms, const Vector &b, const SharedDiagonal &model=SharedDiagonal()) |
| Keys | keys () const |
| std::map< Key, size_t > | getKeyDimMap () const |
| double | error (const VectorValues &x) const |
| double | probPrime (const VectorValues &c) const |
| virtual GaussianFactorGraph | clone () const |
| virtual GaussianFactorGraph::shared_ptr | cloneToPtr () const |
| GaussianFactorGraph | negate () const |
| std::shared_ptr< BayesNetType > | eliminateSequential (OptionalOrderingType orderingType={}, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesNetType > | eliminateSequential (const Ordering &ordering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | eliminateMultifrontal (OptionalOrderingType orderingType={}, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | eliminateMultifrontal (const Ordering &ordering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::pair< std::shared_ptr< BayesNetType >, std::shared_ptr< FactorGraphType > > | eliminatePartialSequential (const Ordering &ordering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::pair< std::shared_ptr< BayesNetType >, std::shared_ptr< FactorGraphType > > | eliminatePartialSequential (const KeyVector &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::pair< std::shared_ptr< BayesTreeType >, std::shared_ptr< FactorGraphType > > | eliminatePartialMultifrontal (const Ordering &ordering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::pair< std::shared_ptr< BayesTreeType >, std::shared_ptr< FactorGraphType > > | eliminatePartialMultifrontal (const KeyVector &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesNetType > | marginalMultifrontalBayesNet (const Ordering &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesNetType > | marginalMultifrontalBayesNet (const KeyVector &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesNetType > | marginalMultifrontalBayesNet (const Ordering &variables, const Ordering &marginalizedVariableOrdering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesNetType > | marginalMultifrontalBayesNet (const KeyVector &variables, const Ordering &marginalizedVariableOrdering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | marginalMultifrontalBayesTree (const Ordering &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | marginalMultifrontalBayesTree (const KeyVector &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | marginalMultifrontalBayesTree (const Ordering &variables, const Ordering &marginalizedVariableOrdering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< BayesTreeType > | marginalMultifrontalBayesTree (const KeyVector &variables, const Ordering &marginalizedVariableOrdering, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
| std::shared_ptr< FactorGraphType > | marginal (const KeyVector &variables, const Eliminate &function=EliminationTraitsType::DefaultEliminate, OptionalVariableIndex variableIndex={}) const |
Constructors | |
| GaussianFactorGraph () | |
| GaussianFactorGraph (std::initializer_list< sharedFactor > factors) | |
| template<typename ITERATOR > | |
| GaussianFactorGraph (ITERATOR firstFactor, ITERATOR lastFactor) | |
| template<class CONTAINER > | |
| GaussianFactorGraph (const CONTAINER &factors) | |
| template<class DERIVEDFACTOR > | |
| GaussianFactorGraph (const FactorGraph< DERIVEDFACTOR > &graph) | |
Testable | |
| bool | equals (const This &fg, double tol=1e-9) const |
Linear Algebra | |
| std::vector< std::tuple< int, int, double > > | sparseJacobian (const Ordering &ordering, size_t &nrows, size_t &ncols) const |
| std::vector< std::tuple< int, int, double > > | sparseJacobian () const |
| Matrix | sparseJacobian_ () const |
| Matrix | augmentedJacobian (const Ordering &ordering) const |
| Matrix | augmentedJacobian () const |
| std::pair< Matrix, Vector > | jacobian (const Ordering &ordering) const |
| std::pair< Matrix, Vector > | jacobian () const |
| Matrix | augmentedHessian (const Ordering &ordering) const |
| Matrix | augmentedHessian () const |
| std::pair< Matrix, Vector > | hessian (const Ordering &ordering) const |
| std::pair< Matrix, Vector > | hessian () const |
| virtual VectorValues | hessianDiagonal () const |
| virtual std::map< Key, Matrix > | hessianBlockDiagonal () const |
| VectorValues | optimize (const Eliminate &function=EliminationTraitsType::DefaultEliminate) const |
| VectorValues | optimize (const Ordering &, const Eliminate &function=EliminationTraitsType::DefaultEliminate) const |
| VectorValues | optimizeDensely () const |
| VectorValues | gradient (const VectorValues &x0) const |
| virtual VectorValues | gradientAtZero () const |
| VectorValues | optimizeGradientSearch () const |
| VectorValues | transposeMultiply (const Errors &e) const |
| void | transposeMultiplyAdd (double alpha, const Errors &e, VectorValues &x) const |
| Errors | gaussianErrors (const VectorValues &x) const |
| Errors | operator* (const VectorValues &x) const |
| void | multiplyHessianAdd (double alpha, const VectorValues &x, VectorValues &y) const |
| void | multiplyInPlace (const VectorValues &x, Errors &e) const |
| void | multiplyInPlace (const VectorValues &x, const Errors::iterator &e) const |
| void | printErrors (const VectorValues &x, const std::string &str="GaussianFactorGraph: ", const KeyFormatter &keyFormatter=DefaultKeyFormatter, const std::function< bool(const Factor *, double, size_t)> &printCondition=[](const Factor *, double, size_t) { return true;}) const |
Adding Single Factors | |
| void | reserve (size_t size) |
| IsDerived< DERIVEDFACTOR > | push_back (std::shared_ptr< DERIVEDFACTOR > factor) |
| Add a factor directly using a shared_ptr. | |
| IsDerived< DERIVEDFACTOR > | push_back (const DERIVEDFACTOR &factor) |
| IsDerived< DERIVEDFACTOR > | emplace_shared (Args &&... args) |
| Emplace a shared pointer to factor of given type. | |
| IsDerived< DERIVEDFACTOR > | add (std::shared_ptr< DERIVEDFACTOR > factor) |
add is a synonym for push_back. | |
Adding via iterators | |
| HasDerivedElementType< ITERATOR > | push_back (ITERATOR firstFactor, ITERATOR lastFactor) |
| HasDerivedValueType< ITERATOR > | push_back (ITERATOR firstFactor, ITERATOR lastFactor) |
| Push back many factors with an iterator (factors are copied) | |
Adding via container | |
| HasDerivedElementType< CONTAINER > | push_back (const CONTAINER &container) |
| HasDerivedValueType< CONTAINER > | push_back (const CONTAINER &container) |
| Push back non-pointer objects in a container (factors are copied). | |
| void | add (const FACTOR_OR_CONTAINER &factorOrContainer) |
Specialized versions | |
| std::enable_if< std::is_base_of< This, typename CLIQUE::FactorGraphType >::value >::type | push_back (const BayesTree< CLIQUE > &bayesTree) |
| FactorIndices | add_factors (const CONTAINER &factors, bool useEmptySlots=false) |
Testable | |
| virtual void | print (const std::string &s="FactorGraph", const KeyFormatter &formatter=DefaultKeyFormatter) const |
| Print out graph to std::cout, with optional key formatter. | |
| bool | equals (const This &fg, double tol=1e-9) const |
| Check equality up to tolerance. | |
Standard Interface | |
| size_t | size () const |
| bool | empty () const |
| const sharedFactor | at (size_t i) const |
| sharedFactor & | at (size_t i) |
| const sharedFactor | operator[] (size_t i) const |
| sharedFactor & | operator[] (size_t i) |
| const_iterator | begin () const |
| const_iterator | end () const |
| sharedFactor | front () const |
| sharedFactor | back () const |
| double | error (const HybridValues &values) const |
Modifying Factor Graphs (imperative, discouraged) | |
| iterator | begin () |
| iterator | end () |
| virtual void | resize (size_t size) |
| void | remove (size_t i) |
| void | replace (size_t index, sharedFactor factor) |
| iterator | erase (iterator item) |
| iterator | erase (iterator first, iterator last) |
Graph Display | |
| void | dot (std::ostream &os, const KeyFormatter &keyFormatter=DefaultKeyFormatter, const DotWriter &writer=DotWriter()) const |
| Output to graphviz format, stream version. | |
| std::string | dot (const KeyFormatter &keyFormatter=DefaultKeyFormatter, const DotWriter &writer=DotWriter()) const |
| Output to graphviz format string. | |
| void | saveGraph (const std::string &filename, const KeyFormatter &keyFormatter=DefaultKeyFormatter, const DotWriter &writer=DotWriter()) const |
| output to file with graphviz format. | |
Advanced Interface | |
| size_t | nrFactors () const |
| KeyVector | keyVector () const |
| bool | exists (size_t idx) const |
Protected Member Functions | |
| bool | isEqual (const FactorGraph &other) const |
| Check exact equality of the factor pointers. Useful for derived ==. | |
Protected Attributes | |
| FastVector< sharedFactor > | factors_ |
Friends | |
| bool | operator== (const GaussianFactorGraph &lhs, const GaussianFactorGraph &rhs) |
| Check exact equality. | |
Detailed Description
A Linear Factor Graph is a factor graph where all factors are Gaussian, i.e. Factor == GaussianFactor VectorValues = A values structure of vectors Most of the time, linear factor graphs arise by linearizing a non-linear factor graph.
Member Typedef Documentation
◆ EliminationResult
|
inherited |
The pair of conditional and remaining factor produced by a single dense elimination step on a subgraph.
◆ Keys
Return the set of variables involved in the factors (computes a set union).
◆ OptionalVariableIndex
|
inherited |
Typedef for an optional variable index as an argument to elimination functions It is an optional to a constant reference
Constructor & Destructor Documentation
◆ GaussianFactorGraph() [1/5]
|
inline |
Default constructor
◆ GaussianFactorGraph() [2/5]
|
inline |
Construct from an initializer lists of GaussianFactor shared pointers. Example: GaussianFactorGraph graph = { factor1, factor2, factor3 };
◆ GaussianFactorGraph() [3/5]
|
inline |
Construct from iterator over factors
◆ GaussianFactorGraph() [4/5]
|
inlineexplicit |
Construct from container of factors (shared_ptr or plain objects)
◆ GaussianFactorGraph() [5/5]
|
inline |
Implicit copy/downcast constructor to override explicit template container constructor
Member Function Documentation
◆ add() [1/8]
|
inline |
Add a factor by value - makes a copy
◆ add() [2/8]
|
inline |
Add a factor by pointer - stores pointer without copying the factor
◆ add() [3/8]
|
inline |
Add a null factor
◆ add() [4/8]
|
inline |
Add a unary factor
◆ add() [5/8]
|
inline |
Add a binary factor
◆ add() [6/8]
|
inline |
Add a ternary factor
◆ add() [7/8]
|
inline |
Add an n-ary factor
◆ add() [8/8]
|
inlineinherited |
Add a factor or container of factors, including STL collections, BayesTrees, etc.
◆ add_factors()
|
inherited |
Add new factors to a factor graph and returns a list of new factor indices, optionally finding and reusing empty factor slots.
◆ at() [1/2]
|
inlineinherited |
Get a specific factor by index (this checks array bounds and may throw an exception, as opposed to operator[] which does not).
◆ at() [2/2]
|
inlineinherited |
Get a specific factor by index (this checks array bounds and may throw an exception, as opposed to operator[] which does not).
◆ augmentedHessian() [1/2]
| Matrix gtsam::GaussianFactorGraph::augmentedHessian | ( | const Ordering & | ordering | ) | const |
Return a dense \( \Lambda \in \mathbb{R}^{n+1 \times n+1} \) Hessian matrix, augmented with the information vector \( \eta \). The augmented Hessian is
\[ \left[ \begin{array}{ccc} \Lambda & \eta \\ \eta^T & c \end{array} \right] \]
and the negative log-likelihood is \( \frac{1}{2} x^T \Lambda x + \eta^T x + c \).
◆ augmentedHessian() [2/2]
| Matrix gtsam::GaussianFactorGraph::augmentedHessian | ( | ) | const |
Return a dense \( \Lambda \in \mathbb{R}^{n+1 \times n+1} \) Hessian matrix, augmented with the information vector \( \eta \). The augmented Hessian is
\[ \left[ \begin{array}{ccc} \Lambda & \eta \\ \eta^T & c \end{array} \right] \]
and the negative log-likelihood is \( \frac{1}{2} x^T \Lambda x + \eta^T x + c \).
◆ augmentedJacobian() [1/2]
| Matrix gtsam::GaussianFactorGraph::augmentedJacobian | ( | const Ordering & | ordering | ) | const |
Return a dense \( [ \;A\;b\; ] \in \mathbb{R}^{m \times n+1} \) Jacobian matrix, augmented with b with the noise models baked into A and b. The negative log-likelihood is \( \frac{1}{2} \Vert Ax-b \Vert^2 \). See also GaussianFactorGraph::jacobian and GaussianFactorGraph::sparseJacobian.
◆ augmentedJacobian() [2/2]
| Matrix gtsam::GaussianFactorGraph::augmentedJacobian | ( | ) | const |
Return a dense \( [ \;A\;b\; ] \in \mathbb{R}^{m \times n+1} \) Jacobian matrix, augmented with b with the noise models baked into A and b. The negative log-likelihood is \( \frac{1}{2} \Vert Ax-b \Vert^2 \). See also GaussianFactorGraph::jacobian and GaussianFactorGraph::sparseJacobian.
◆ back()
|
inlineinherited |
Get the last factor
◆ begin() [1/2]
|
inlineinherited |
Iterator to beginning of factors.
◆ begin() [2/2]
|
inlineinherited |
non-const STL-style begin()
◆ clone()
|
virtual |
Clone() performs a deep-copy of the graph, including all of the factors. Cloning preserves null factors so indices for the original graph are still valid for the cloned graph.
◆ cloneToPtr()
|
virtual |
CloneToPtr() performs a simple assignment to a new graph and returns it. There is no preservation of null factors!
◆ eliminateMultifrontal() [1/2]
|
inherited |
Do multifrontal elimination of all variables to produce a Bayes tree. If an ordering is not provided, the ordering will be computed using either COLAMD or METIS, depending on the parameter orderingType (Ordering::COLAMD or Ordering::METIS)
Example - Full Cholesky elimination in COLAMD order:
Example - Reusing an existing VariableIndex to improve performance, and using COLAMD ordering:
◆ eliminateMultifrontal() [2/2]
|
inherited |
Do multifrontal elimination of all variables to produce a Bayes tree. If an ordering is not provided, the ordering will be computed using either COLAMD or METIS, depending on the parameter orderingType (Ordering::COLAMD or Ordering::METIS)
Example - Full QR elimination in specified order:
◆ eliminatePartialMultifrontal() [1/2]
|
inherited |
Do multifrontal elimination of some variables, in ordering provided, to produce a Bayes tree and a remaining factor graph. This computes the factorization \( p(X) = p(A|B) p(B) \), where \( A = \) variables, \( X \) is all the variables in the factor graph, and \( B = X\backslash A \).
◆ eliminatePartialMultifrontal() [2/2]
|
inherited |
Do multifrontal elimination of the given variables in an ordering computed by COLAMD to produce a Bayes tree and a remaining factor graph. This computes the factorization \( p(X) = p(A|B) p(B) \), where \( A = \) variables, \( X \) is all the variables in the factor graph, and \( B = X\backslash A \).
◆ eliminatePartialSequential() [1/2]
|
inherited |
Do sequential elimination of some variables, in ordering provided, to produce a Bayes net and a remaining factor graph. This computes the factorization \( p(X) = p(A|B) p(B) \), where \( A = \) variables, \( X \) is all the variables in the factor graph, and \( B = X\backslash A \).
◆ eliminatePartialSequential() [2/2]
|
inherited |
Do sequential elimination of the given variables in an ordering computed by COLAMD to produce a Bayes net and a remaining factor graph. This computes the factorization \( p(X) = p(A|B) p(B) \), where \( A = \) variables, \( X \) is all the variables in the factor graph, and \( B = X\backslash A \).
◆ eliminateSequential() [1/2]
|
inherited |
Do sequential elimination of all variables to produce a Bayes net. If an ordering is not provided, the ordering provided by COLAMD will be used.
Example - Full Cholesky elimination in COLAMD order:
Example - METIS ordering for elimination
Example - Reusing an existing VariableIndex to improve performance, and using COLAMD ordering:
◆ eliminateSequential() [2/2]
|
inherited |
Do sequential elimination of all variables to produce a Bayes net.
Example - Full QR elimination in specified order:
Example - Reusing an existing VariableIndex to improve performance:
◆ empty()
|
inlineinherited |
Check if the graph is empty (null factors set by remove() will cause this to return false).
◆ end() [1/2]
|
inlineinherited |
Iterator to end of factors.
◆ end() [2/2]
|
inlineinherited |
non-const STL-style end()
◆ erase() [1/2]
|
inlineinherited |
Erase factor and rearrange other factors to take up the empty space
◆ erase() [2/2]
|
inlineinherited |
Erase factors and rearrange other factors to take up the empty space
◆ error() [1/2]
| double gtsam::GaussianFactorGraph::error | ( | const VectorValues & | x | ) | const |
unnormalized error
◆ error() [2/2]
|
inherited |
Add error for all factors.
◆ exists()
|
inlineinherited |
MATLAB interface utility: Checks whether a factor index idx exists in the graph and is a live pointer
◆ front()
|
inlineinherited |
Get the first factor
◆ gaussianErrors()
| Errors gtsam::GaussianFactorGraph::gaussianErrors | ( | const VectorValues & | x | ) | const |
return A*x-b
◆ gradient()
| VectorValues gtsam::GaussianFactorGraph::gradient | ( | const VectorValues & | x0 | ) | const |
Compute the gradient of the energy function, \( \nabla_{x=x_0} \left\Vert \Sigma^{-1} A x - b \right\Vert^2 \), centered around \( x = x_0 \). The gradient is \( A^T(Ax-b) \).
- Parameters
-
fg The Jacobian factor graph $(A,b)$ x0 The center about which to compute the gradient
- Returns
- The gradient as a VectorValues
◆ gradientAtZero()
|
virtual |
Compute the gradient of the energy function, \( \nabla_{x=0} \left\Vert \Sigma^{-1} A x - b \right\Vert^2 \), centered around zero. The gradient is \( A^T(Ax-b) \).
- Parameters
-
fg The Jacobian factor graph $(A,b)$ [output] g A VectorValues to store the gradient, which must be preallocated, see allocateVectorValues
- Returns
- The gradient as a VectorValues
◆ hessian() [1/2]
| std::pair<Matrix,Vector> gtsam::GaussianFactorGraph::hessian | ( | const Ordering & | ordering | ) | const |
Return the dense Hessian \( \Lambda \) and information vector \( \eta \), with the noise models baked in. The negative log-likelihood is {1}{2} x^T x + ^T x + c. See also GaussianFactorGraph::augmentedHessian.
◆ hessian() [2/2]
| std::pair<Matrix,Vector> gtsam::GaussianFactorGraph::hessian | ( | ) | const |
Return the dense Hessian \( \Lambda \) and information vector \( \eta \), with the noise models baked in. The negative log-likelihood is {1}{2} x^T x + ^T x + c. See also GaussianFactorGraph::augmentedHessian.
◆ hessianBlockDiagonal()
|
virtual |
Return the block diagonal of the Hessian for this factor
◆ hessianDiagonal()
|
virtual |
Return only the diagonal of the Hessian A'*A, as a VectorValues
◆ jacobian() [1/2]
| std::pair<Matrix,Vector> gtsam::GaussianFactorGraph::jacobian | ( | const Ordering & | ordering | ) | const |
Return the dense Jacobian \( A \) and right-hand-side \( b \), with the noise models baked into A and b. The negative log-likelihood is \( \frac{1}{2} \Vert Ax-b \Vert^2 \). See also GaussianFactorGraph::augmentedJacobian and GaussianFactorGraph::sparseJacobian.
◆ jacobian() [2/2]
| std::pair<Matrix,Vector> gtsam::GaussianFactorGraph::jacobian | ( | ) | const |
Return the dense Jacobian \( A \) and right-hand-side \( b \), with the noise models baked into A and b. The negative log-likelihood is \( \frac{1}{2} \Vert Ax-b \Vert^2 \). See also GaussianFactorGraph::augmentedJacobian and GaussianFactorGraph::sparseJacobian.
◆ keyVector()
|
inherited |
Potentially slow function to return all keys involved, sorted, as a vector
◆ marginal()
|
inherited |
Compute the marginal factor graph of the requested variables.
◆ marginalMultifrontalBayesNet() [1/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes net. Uses COLAMD marginalization ordering by default
- Parameters
-
variables Determines the ordered variables whose marginal to compute, will be ordered in the returned BayesNet as specified. function Optional dense elimination function. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesNet() [2/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes net. Uses COLAMD marginalization ordering by default
- Parameters
-
variables Determines the variables whose marginal to compute, will be ordered using COLAMD; use Ordering(variables)to specify the variable ordering.function Optional dense elimination function. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesNet() [3/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes net.
- Parameters
-
variables Determines the ordered variables whose marginal to compute, will be ordered in the returned BayesNet as specified. marginalizedVariableOrdering Ordering for the variables being marginalized out, i.e. all variables not in variables.function Optional dense elimination function. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesNet() [4/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes net.
- Parameters
-
variables Determines the variables whose marginal to compute, will be ordered using COLAMD; use Ordering(variables)to specify the variable ordering.marginalizedVariableOrdering Ordering for the variables being marginalized out, i.e. all variables not in variables.function Optional dense elimination function. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesTree() [1/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes tree. Uses COLAMD marginalization order by default
- Parameters
-
variables Determines the ordered variables whose marginal to compute, will be ordered in the returned BayesNet as specified. function Optional dense elimination function.. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesTree() [2/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes tree. Uses COLAMD marginalization order by default
- Parameters
-
variables Determines the variables whose marginal to compute, will be ordered using COLAMD; use Ordering(variables)to specify the variable ordering.function Optional dense elimination function.. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesTree() [3/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes tree.
- Parameters
-
variables Determines the ordered variables whose marginal to compute, will be ordered in the returned BayesNet as specified. marginalizedVariableOrdering Ordering for the variables being marginalized out, i.e. all variables not in variables.function Optional dense elimination function.. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ marginalMultifrontalBayesTree() [4/4]
|
inherited |
Compute the marginal of the requested variables and return the result as a Bayes tree.
- Parameters
-
variables Determines the variables whose marginal to compute, will be ordered using COLAMD; use Ordering(variables)to specify the variable ordering.marginalizedVariableOrdering Ordering for the variables being marginalized out, i.e. all variables not in variables.function Optional dense elimination function.. variableIndex Optional pre-computed VariableIndex for the factor graph, if not provided one will be computed.
◆ multiplyHessianAdd()
| void gtsam::GaussianFactorGraph::multiplyHessianAdd | ( | double | alpha, |
| const VectorValues & | x, | ||
| VectorValues & | y | ||
| ) | const |
y += alpha*A'A*x
◆ multiplyInPlace() [1/2]
| void gtsam::GaussianFactorGraph::multiplyInPlace | ( | const VectorValues & | x, |
| Errors & | e | ||
| ) | const |
In-place version e <- A*x that overwrites e.
◆ multiplyInPlace() [2/2]
| void gtsam::GaussianFactorGraph::multiplyInPlace | ( | const VectorValues & | x, |
| const Errors::iterator & | e | ||
| ) | const |
In-place version e <- A*x that takes an iterator.
◆ negate()
| GaussianFactorGraph gtsam::GaussianFactorGraph::negate | ( | ) | const |
Returns the negation of all factors in this graph - corresponds to antifactors. Will convert all factors to HessianFactors due to negation of information. Cloning preserves null factors so indices for the original graph are still valid for the cloned graph.
◆ nrFactors()
|
inherited |
return the number of non-null factors
◆ operator*()
| Errors gtsam::GaussianFactorGraph::operator* | ( | const VectorValues & | x | ) | const |
return A*x
◆ operator[]() [1/2]
|
inlineinherited |
Get a specific factor by index (this does not check array bounds, as opposed to at() which does).
◆ operator[]() [2/2]
|
inlineinherited |
Get a specific factor by index (this does not check array bounds, as opposed to at() which does).
◆ optimize() [1/2]
| VectorValues gtsam::GaussianFactorGraph::optimize | ( | const Eliminate & | function = EliminationTraitsType::DefaultEliminate | ) | const |
Solve the factor graph by performing multifrontal variable elimination in COLAMD order using the dense elimination function specified in function (default EliminatePreferCholesky), followed by back-substitution in the Bayes tree resulting from elimination. Is equivalent to calling graph.eliminateMultifrontal()->optimize().
◆ optimize() [2/2]
| VectorValues gtsam::GaussianFactorGraph::optimize | ( | const Ordering & | , |
| const Eliminate & | function = EliminationTraitsType::DefaultEliminate |
||
| ) | const |
Solve the factor graph by performing multifrontal variable elimination in COLAMD order using the dense elimination function specified in function (default EliminatePreferCholesky), followed by back-substitution in the Bayes tree resulting from elimination. Is equivalent to calling graph.eliminateMultifrontal()->optimize().
◆ optimizeDensely()
| VectorValues gtsam::GaussianFactorGraph::optimizeDensely | ( | ) | const |
Optimize using Eigen's dense Cholesky factorization
◆ optimizeGradientSearch()
| VectorValues gtsam::GaussianFactorGraph::optimizeGradientSearch | ( | ) | const |
Optimize along the gradient direction, with a closed-form computation to perform the line search. The gradient is computed about \( \delta x=0 \).
This function returns \( \delta x \) that minimizes a reparametrized problem. The error function of a GaussianBayesNet is
\[ f(\delta x) = \frac{1}{2} |R \delta x - d|^2 = \frac{1}{2}d^T d - d^T R \delta x + \frac{1}{2} \delta x^T R^T R \delta x \]
with gradient and Hessian
\[ g(\delta x) = R^T(R\delta x - d), \qquad G(\delta x) = R^T R. \]
This function performs the line search in the direction of the gradient evaluated at \( g = g(\delta x = 0) \) with step size \( \alpha \) that minimizes \( f(\delta x = \alpha g) \):
\[ f(\alpha) = \frac{1}{2} d^T d + g^T \delta x + \frac{1}{2} \alpha^2 g^T G g \]
Optimizing by setting the derivative to zero yields \( \hat \alpha = (-g^T g) / (g^T G g) \). For efficiency, this function evaluates the denominator without computing the Hessian \( G \), returning
\[ \delta x = \hat\alpha g = \frac{-g^T g}{(R g)^T(R g)} \]
◆ probPrime()
| double gtsam::GaussianFactorGraph::probPrime | ( | const VectorValues & | c | ) | const |
Unnormalized probability. O(n)
◆ push_back() [1/4]
|
inlineinherited |
Add a factor by value, will be copy-constructed (use push_back with a shared_ptr to avoid the copy).
◆ push_back() [2/4]
|
inlineinherited |
Push back many factors with an iterator over shared_ptr (factors are not copied)
◆ push_back() [3/4]
|
inlineinherited |
Push back many factors as shared_ptr's in a container (factors are not copied)
◆ push_back() [4/4]
|
inlineinherited |
Push back a BayesTree as a collection of factors. NOTE: This should be hidden in derived classes in favor of a type-specialized version that calls this templated function.
◆ remove()
|
inlineinherited |
delete factor without re-arranging indexes by inserting a nullptr pointer
◆ replace()
|
inlineinherited |
replace a factor by index
◆ reserve()
|
inlineinherited |
Reserve space for the specified number of factors if you know in advance how many there will be (works like FastVector::reserve).
◆ resize()
|
inlinevirtualinherited |
Directly resize the number of factors in the graph. If the new size is less than the original, factors at the end will be removed. If the new size is larger than the original, null factors will be appended.
◆ size()
|
inlineinherited |
return the number of factors (including any null factors set by remove() ).
◆ sparseJacobian() [1/2]
| std::vector<std::tuple<int, int, double> > gtsam::GaussianFactorGraph::sparseJacobian | ( | const Ordering & | ordering, |
| size_t & | nrows, | ||
| size_t & | ncols | ||
| ) | const |
Returns a sparse augmented Jacbian matrix as a vector of i, j, and s, where i(k) and j(k) are the base 0 row and column indices, and s(k) is the entry as a double. The standard deviations are baked into A and b
- Returns
- the sparse matrix as a std::vector of std::tuples
- Parameters
-
ordering the column ordering [out] nrows The number of rows in the augmented Jacobian [out] ncols The number of columns in the augmented Jacobian
◆ sparseJacobian() [2/2]
| std::vector<std::tuple<int, int, double> > gtsam::GaussianFactorGraph::sparseJacobian | ( | ) | const |
Returns a sparse augmented Jacobian matrix with default ordering
◆ sparseJacobian_()
| Matrix gtsam::GaussianFactorGraph::sparseJacobian_ | ( | ) | const |
Matrix version of sparseJacobian: generates a 3*m matrix with [i,j,s] entries such that S(i(k),j(k)) = s(k), which can be given to MATLAB's sparse. Note: i, j are 1-indexed. The standard deviations are baked into A and b
◆ transposeMultiply()
| VectorValues gtsam::GaussianFactorGraph::transposeMultiply | ( | const Errors & | e | ) | const |
x = A'*e
◆ transposeMultiplyAdd()
| void gtsam::GaussianFactorGraph::transposeMultiplyAdd | ( | double | alpha, |
| const Errors & | e, | ||
| VectorValues & | x | ||
| ) | const |
x += alpha*A'*e
Member Data Documentation
◆ factors_
|
protectedinherited |
concept check, makes sure FACTOR defines print and equals Collection of factors
The documentation for this class was generated from the following file:
- /home/docs/checkouts/readthedocs.org/user_builds/gtsam-jlblanco-docs/checkouts/latest/gtsam/linear/GaussianFactorGraph.h
 1.8.13
1.8.13