
Public Types | |
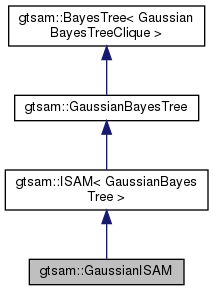
| typedef ISAM< GaussianBayesTree > | Base |
| typedef GaussianISAM | This |
| typedef std::shared_ptr< This > | shared_ptr |
| typedef Base::BayesNetType | BayesNetType |
| typedef Base::FactorGraphType | FactorGraphType |
| typedef Base::Clique | Clique |
| typedef Base::sharedClique | sharedClique |
| typedef Base::Cliques | Cliques |
| typedef Clique | Node |
| Synonym for Clique (TODO: remove) | |
| typedef sharedClique | sharedNode |
| Synonym for sharedClique (TODO: remove) | |
| typedef GaussianBayesTreeClique ::ConditionalType | ConditionalType |
| typedef std::shared_ptr< ConditionalType > | sharedConditional |
| typedef std::shared_ptr< BayesNetType > | sharedBayesNet |
| typedef GaussianBayesTreeClique ::FactorType | FactorType |
| typedef std::shared_ptr< FactorType > | sharedFactor |
| typedef std::shared_ptr< FactorGraphType > | sharedFactorGraph |
| typedef ConcurrentMap< Key, sharedClique > | Nodes |
| typedef FastVector< sharedClique > | Roots |
Public Member Functions | |
| bool | equals (const This &other, double tol=1e-9) const |
| VectorValues | optimize () const |
| VectorValues | optimizeGradientSearch () const |
| VectorValues | gradient (const VectorValues &x0) const |
| VectorValues | gradientAtZero () const |
| double | error (const VectorValues &x) const |
| double | determinant () const |
| double | logDeterminant () const |
| Matrix | marginalCovariance (Key key) const |
Standard Constructors | |
| GaussianISAM () | |
| GaussianISAM (const GaussianBayesTree &bayesTree) | |
Advanced Interface Interface | |
| void | update (const FactorGraphType &newFactors, const Eliminate &function=EliminationTraitsType::DefaultEliminate) |
| void | updateInternal (const FactorGraphType &newFactors, Cliques *orphans, const Eliminate &function=EliminationTraitsType::DefaultEliminate) |
Standard Interface | |
| size_t | size () const |
| bool | empty () const |
| const Nodes & | nodes () const |
| sharedClique | operator[] (Key j) const |
| const Roots & | roots () const |
| const sharedClique & | clique (Key j) const |
| BayesTreeCliqueData | getCliqueData () const |
| size_t | numCachedSeparatorMarginals () const |
| sharedConditional | marginalFactor (Key j, const Eliminate &function=EliminationTraitsType::DefaultEliminate) const |
| sharedFactorGraph | joint (Key j1, Key j2, const Eliminate &function=EliminationTraitsType::DefaultEliminate) const |
| sharedBayesNet | jointBayesNet (Key j1, Key j2, const Eliminate &function=EliminationTraitsType::DefaultEliminate) const |
Graph Display | |
| void | dot (std::ostream &os, const KeyFormatter &keyFormatter=DefaultKeyFormatter) const |
| Output to graphviz format, stream version. | |
| std::string | dot (const KeyFormatter &keyFormatter=DefaultKeyFormatter) const |
| Output to graphviz format string. | |
| void | saveGraph (const std::string &filename, const KeyFormatter &keyFormatter=DefaultKeyFormatter) const |
| output to file with graphviz format. | |
Protected Attributes | |
| Nodes | nodes_ |
| Roots | roots_ |
Testable | |
| void | print (const std::string &s="", const KeyFormatter &keyFormatter=DefaultKeyFormatter) const |
| bool | equals (const This &other, double tol=1e-9) const |
Advanced Interface | |
| Key | findParentClique (const CONTAINER &parents) const |
| void | clear () |
| void | deleteCachedShortcuts () |
| void | removePath (sharedClique clique, BayesNetType *bn, Cliques *orphans) |
| void | removeTop (const KeyVector &keys, BayesNetType *bn, Cliques *orphans) |
| Cliques | removeSubtree (const sharedClique &subtree) |
| void | insertRoot (const sharedClique &subtree) |
| void | addClique (const sharedClique &clique, const sharedClique &parent_clique=sharedClique()) |
| void | addFactorsToGraph (FactorGraph< FactorType > *graph) const |
| void | getCliqueData (sharedClique clique, BayesTreeCliqueData *stats) const |
| void | dot (std::ostream &s, sharedClique clique, const KeyFormatter &keyFormatter, int parentnum=0) const |
| void | removeClique (sharedClique clique) |
| void | fillNodesIndex (const sharedClique &subtree) |
Member Typedef Documentation
◆ Nodes
|
inherited |
Map from keys to Clique
◆ Roots
|
inherited |
Root cliques
Constructor & Destructor Documentation
◆ GaussianISAM() [1/2]
| gtsam::GaussianISAM::GaussianISAM | ( | ) |
Create an empty Bayes Tree
◆ GaussianISAM() [2/2]
| gtsam::GaussianISAM::GaussianISAM | ( | const GaussianBayesTree & | bayesTree | ) |
Copy constructor
Member Function Documentation
◆ addClique()
|
inherited |
add a clique (top down)
◆ addFactorsToGraph()
|
inherited |
Add all cliques in this BayesTree to the specified factor graph
◆ clear()
|
inherited |
Remove all nodes
◆ clique()
|
inlineinherited |
alternate syntax for matlab: find the clique that contains the variable with Key j
◆ deleteCachedShortcuts()
|
inherited |
Clear all shortcut caches - use before timing on marginal calculation to avoid residual cache data
◆ determinant()
|
inherited |
Computes the determinant of a GassianBayesTree, as if the Bayes tree is reorganized into a matrix. A GassianBayesTree is equivalent to an upper triangular matrix, and for an upper triangular matrix determinant is the product of the diagonal elements. Instead of actually multiplying we add the logarithms of the diagonal elements and take the exponent at the end because this is more numerically stable.
◆ dot()
|
protectedinherited |
private helper method for saving the Tree to a text file in GraphViz format
◆ empty()
|
inlineinherited |
Check if there are any cliques in the tree
◆ equals() [1/2]
|
inherited |
Check equality
◆ equals() [2/2]
|
protectedinherited |
check equality
◆ error()
|
inherited |
0.5 * sum of squared Mahalanobis distances.
◆ fillNodesIndex()
|
protectedinherited |
Fill the nodes index for a subtree
◆ findParentClique()
|
inherited |
Find parent clique of a conditional. It will look at all parents and return the one with the lowest index in the ordering.
◆ getCliqueData() [1/2]
|
inherited |
Gather data on all cliques
◆ getCliqueData() [2/2]
|
protectedinherited |
Gather data on a single clique
◆ gradient()
|
inherited |
Compute the gradient of the energy function, \( \nabla_{x=x_0} \left\Vert \Sigma^{-1} R x - d \right\Vert^2 \), centered around \( x = x_0 \). The gradient is \( R^T(Rx-d) \).
- Parameters
-
x0 The center about which to compute the gradient
- Returns
- The gradient as a VectorValues
◆ gradientAtZero()
|
inherited |
Compute the gradient of the energy function, \( \nabla_{x=0} \left\Vert \Sigma^{-1} R x - d \right\Vert^2 \), centered around zero. The gradient about zero is \( -R^T d \). See also gradient(const GaussianBayesNet&, const VectorValues&).
- Returns
- A VectorValues storing the gradient.
◆ insertRoot()
|
inherited |
Insert a new subtree with known parent clique. This function does not check that the specified parent is the correct parent. This function updates all of the internal data structures associated with adding a subtree, such as populating the nodes index.
◆ joint()
|
inherited |
return joint on two variables Limitation: can only calculate joint if cliques are disjoint or one of them is root
◆ jointBayesNet()
|
inherited |
return joint on two variables as a BayesNet Limitation: can only calculate joint if cliques are disjoint or one of them is root
◆ logDeterminant()
|
inherited |
Computes the determinant of a GassianBayesTree, as if the Bayes tree is reorganized into a matrix. A GassianBayesTree is equivalent to an upper triangular matrix, and for an upper triangular matrix determinant is the product of the diagonal elements. Instead of actually multiplying we add the logarithms of the diagonal elements and take the exponent at the end because this is more numerically stable.
◆ marginalCovariance()
|
inherited |
Return the marginal on the requested variable as a covariance matrix. See also marginalFactor().
◆ marginalFactor()
|
inherited |
Return marginal on any variable. Note that this actually returns a conditional, for which a solution may be directly obtained by calling .solve() on the returned object. Alternatively, it may be directly used as its factor base class. For example, for Gaussian systems, this returns a GaussianConditional, which inherits from JacobianFactor and GaussianFactor.
◆ nodes()
|
inlineinherited |
Return nodes. Each node is a clique of variables obtained after elimination.
◆ numCachedSeparatorMarginals()
|
inherited |
Collect number of cliques with cached separator marginals
◆ operator[]()
|
inlineinherited |
Access node by variable
◆ optimize()
|
inherited |
Recursively optimize the BayesTree to produce a vector solution.
◆ optimizeGradientSearch()
|
inherited |
Optimize along the gradient direction, with a closed-form computation to perform the line search. The gradient is computed about \( \delta x=0 \).
This function returns \( \delta x \) that minimizes a reparametrized problem. The error function of a GaussianBayesNet is
\[ f(\delta x) = \frac{1}{2} |R \delta x - d|^2 = \frac{1}{2}d^T d - d^T R \delta x + \frac{1}{2} \delta x^T R^T R \delta x \]
with gradient and Hessian
\[ g(\delta x) = R^T(R\delta x - d), \qquad G(\delta x) = R^T R. \]
This function performs the line search in the direction of the gradient evaluated at \( g = g(\delta x = 0) \) with step size \( \alpha \) that minimizes \( f(\delta x = \alpha g) \):
\[ f(\alpha) = \frac{1}{2} d^T d + g^T \delta x + \frac{1}{2} \alpha^2 g^T G g \]
Optimizing by setting the derivative to zero yields \( \hat \alpha = (-g^T g) / (g^T G g) \). For efficiency, this function evaluates the denominator without computing the Hessian \( G \), returning
\[ \delta x = \hat\alpha g = \frac{-g^T g}{(R g)^T(R g)} \]
◆ print()
|
inherited |
◆ removeClique()
|
protectedinherited |
remove a clique: warning, can result in a forest
◆ removePath()
|
inherited |
Remove path from clique to root and return that path as factors plus a list of orphaned subtree roots. Used in removeTop below.
◆ removeSubtree()
|
inherited |
Remove the requested subtree.
◆ removeTop()
|
inherited |
Given a list of indices, turn "contaminated" part of the tree back into a factor graph. Factors and orphans are added to the in/out arguments.
◆ roots()
|
inlineinherited |
return root cliques
◆ size()
|
inherited |
number of cliques
◆ update()
|
inherited |
update the Bayes tree with a set of new factors, typically derived from measurements
- Parameters
-
newFactors is a factor graph that contains the new factors function an elimination routine
◆ updateInternal()
|
inherited |
updateInternal provides access to list of orphans for drawing purposes
Member Data Documentation
◆ nodes_
|
protectedinherited |
Map from indices to Clique
◆ roots_
|
protectedinherited |
Root cliques
The documentation for this class was generated from the following file:
- /home/docs/checkouts/readthedocs.org/user_builds/gtsam-jlblanco-docs/checkouts/latest/gtsam/linear/GaussianISAM.h
 1.8.13
1.8.13